Goodness-of-fit measures to compare observed and simulated time series with hydroGOF
Mauricio Zambrano-Bigiarini1
version 0.4, 30-Apr-2026
Source:vignettes/hydroGOF_Vignette.Rmd
hydroGOF_Vignette.RmdCitation
If you use hydroGOF, please cite it as Zambrano-Bigiarini (2026):
Zambrano-Bigiarini, M. (2026) hydroGOF: Goodness-of-fit functions for comparison of simulated and observed hydrological time series R package version 0.7-0. URL: https://cran.r-project.org/package=hydroGOF. doi:10.32614/CRAN.package.hydroGOF.
Installation
Installing the latest stable version (from CRAN):
install.packages("hydroGOF")Alternatively, you can also try the under-development version (from Github):
if (!require(devtools)) install.packages("devtools")
library(devtools)
install_github("hzambran/hydroGOF")Setting up the environment
Loading the hydroGOF package, which contains data and functions used in this analysis:
## Loading required package: zoo##
## Attaching package: 'zoo'## The following objects are masked from 'package:base':
##
## as.Date, as.Date.numericExample using NSE
The following examples use the well-known Nash-Sutcliffe efficiency (NSE), but you can repeat the computations using any of the goodness-of-fit measures included in the hydroGOF package (e.g., KGE, ubRMSE, dr).
Ex 1: ideal integer values
Basic ideal case with a numeric sequence of integers:
obs <- 1:10
sim <- 1:10
NSE(sim, obs)## [1] 1
obs <- 1:10
sim <- 2:11
NSE(sim, obs)## [1] 0.8787879Ex 2: ideal real values
Basic ideal case with a numeric sequence of integers:
obs <- 1:10 + 0.1
sim <- 1:10 +0.1
NSE(sim, obs)## [1] 1
obs <- 1:10 + 0.1
sim <- 2:11 + 0.1
NSE(sim, obs)## [1] 0.8787879Ex 3: ideal daily ts
From this example onwards, a streamflow time series will be used.
First, we load the daily streamflows of the Ega River (Spain), from 1961 to 1970:
data(EgaEnEstellaQts)
obs <- EgaEnEstellaQtsGenerating a simulated daily time series, initially equal to the observed series:
sim <- obs Visualising the observed and simulated time series:
ggof(sim, obs)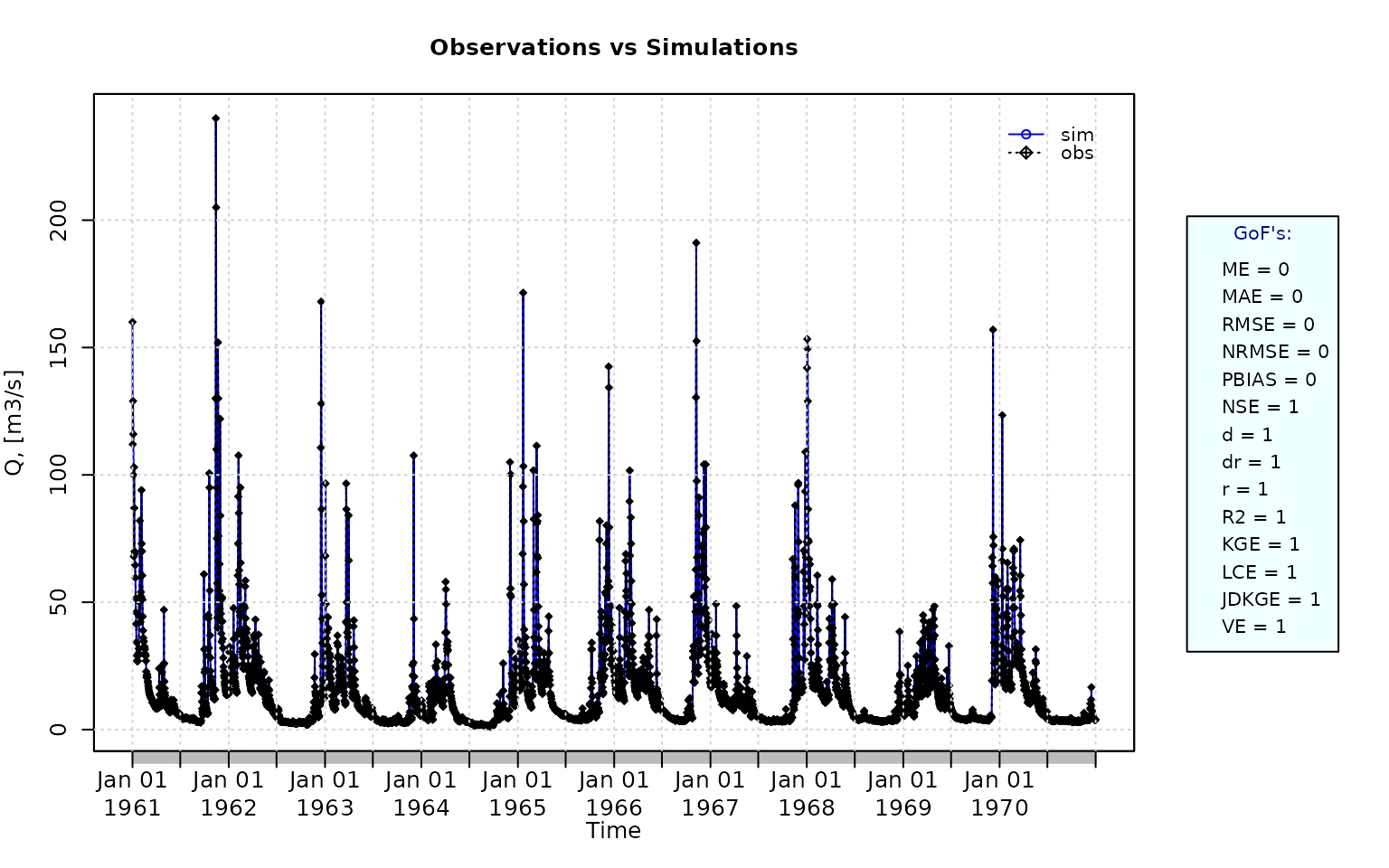
Computing the ‘NSE’ for the “best” (unattainable) case
NSE(sim=sim, obs=obs)## [1] 1Ex 4: random noise in the first half of the simulated values
NSE for simulated values equal to observations plus random noise on the first half of the observed values.
This random noise has more relative importance for low flows than for medium and high flows.
Randomly changing the first 1826 elements of ‘sim’, by using a normal distribution with mean 10 and standard deviation equal to 1 (default of ‘rnorm’).
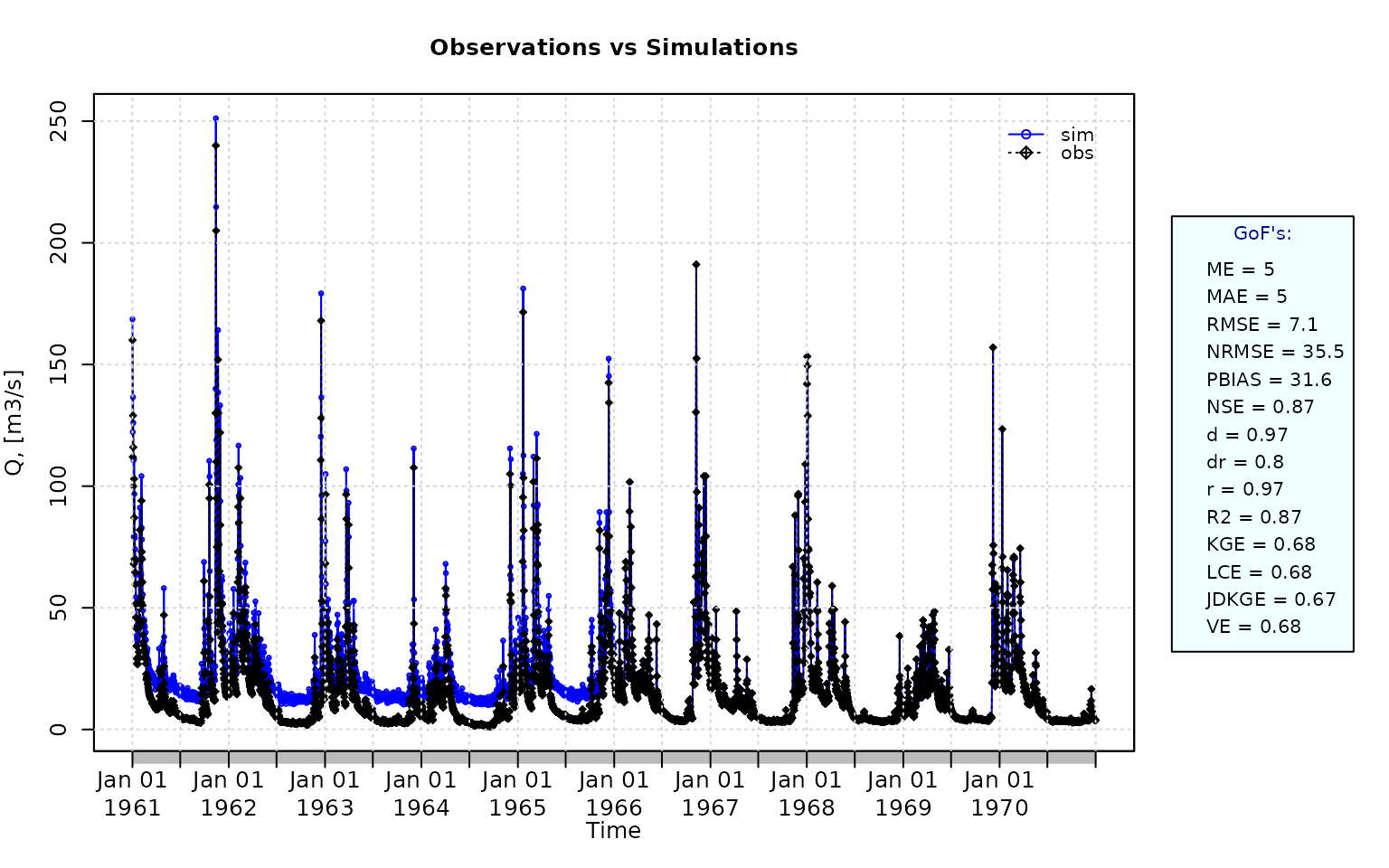
NSE(sim=sim, obs=obs)## [1] 0.8739885Let’s have a look at other goodness-of-fit measures:
mNSE(sim=sim, obs=obs) # Modified NSE## [1] 0.6049584
rNSE(sim=sim, obs=obs) # Relative NSE## [1] -0.5687206
wNSE(sim=sim, obs=obs) # Weighted NSE## [1] 0.9769178
wsNSE(sim=sim, obs=obs) # Weighted Seasonal NSE## [1] 0.9686072
KGE(sim=sim, obs=obs) # Kling-Gupta efficiency (KGE), 2009## [1] 0.6805856
KGE(sim=sim, obs=obs, method="2012") # Kling-Gupta efficiency (KGE), 2012## [1] 0.61689
KGE(sim=sim, obs=obs, method="2021") # Kling-Gupta efficiency (KGE), 2021## [1] 0.7461172
KGElf(sim=sim, obs=obs) # KGE for low flows## [1] 0.5170138
KGEnp(sim=sim, obs=obs) # Non-parametric KGE## [1] 0.6340134
sKGE(sim=sim, obs=obs) # Split KGE## [1] 0.6547522
KGEkm(sim=sim, obs=obs) # Knowable Moments KGE## [1] 0.6467934
JDKGE(sim=sim, obs=obs) # Joint Divergence KGE## [1] 0.6080188
d(sim=sim, obs=obs) # Index of Agreement## [1] 0.9697286
dr(sim=sim, obs=obs) # Refined Index of Agreement## [1] 0.8024792
md(sim=sim, obs=obs) # Modified Index of Agreement## [1] 0.7980307
rd(sim=sim, obs=obs) # Relative Index of Agreement## [1] 0.6231506
VE(sim=sim, obs=obs) # Volumetric Efficiency## [1] 0.6838531
cp(sim=sim, obs=obs) # Coefficient of Persistence## [1] 0.4683536
APFB(sim=sim, obs=obs) # Annual Peak Flow Bias## [1] 0.03202514
HFB(sim=sim, obs=obs) # High Flow Bias## [1] 0.08706977
LME(sim=sim, obs=obs) # Liu-Mean Efficiency## [1] 0.6838398
LCE(sim=sim, obs=obs) # Lee and Choi Efficiency## [1] 0.6777786
PMR(sim=sim, obs=obs) # Proxy for Model Robustness## [1] 0.3156247
pbias(sim=sim, obs=obs) # Percent bias (PBIAS)## [1] 31.6
pbiasfdc(sim=sim, obs=obs) # PBIAS in the slope of the midsegment of the FDC## [Note: 'thr.shw' was set to FALSE to avoid confusing legends...]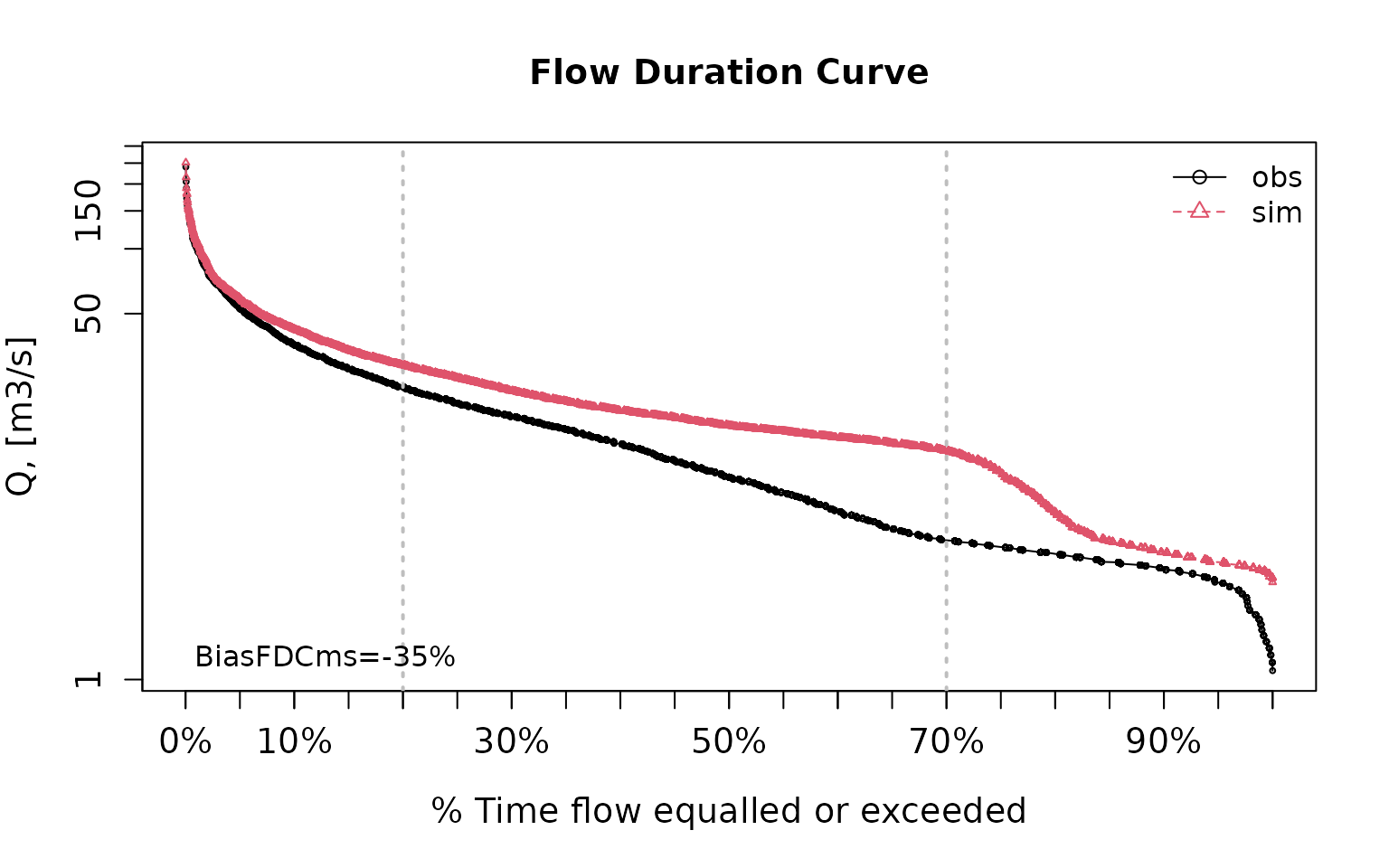
## [1] -34.95419
me(sim=sim, obs=obs) # Mean Error## [1] 4.998998
mae(sim=sim, obs=obs) # Mean Absolute Error## [1] 4.998998
mse(sim=sim, obs=obs) # Mean Squared Error ## [1] 50.46771
rmse(sim=sim, obs=obs) # Root Mean Square Error (RMSE)## [1] 7.104063
ubRMSE(sim=sim, obs=obs) # Unbiased RMSE## [1] 5.047547
nrmse(sim=sim, obs=obs, norm="sd") # Normalised Root Mean Square Error## [1] 35.5
nrmse(sim=sim, obs=obs, norm="maxmin") # Normalised Root Mean Square Error## [1] 3
nrmse(sim=sim, obs=obs, norm="mean") # Normalised Root Mean Square Error## [1] 44.9
nrmse(sim=sim, obs=obs, norm="IQR") # Normalised Root Mean Square Error## [1] 47.1
rPearson(sim=sim, obs=obs) # Pearson correlation coefficient## [1] 0.9698058
rSpearman(sim=sim, obs=obs) # Spearman rank correlation coefficient## [1] 0.8362479
R2(sim=sim, obs=obs) # Coefficient of determination (R2)## [1] 0.8739885
br2(sim=sim, obs=obs) # R2 multiplied by the slope of the regression line## [1] 0.7780545Ex 5: random noise and logarithmic transformation
NSE for simulated values equal to observations plus random noise on the first half of the observed values and applying (natural) logarithm to obs against sim during computations.
NSE(sim=sim, obs=obs, fun=log)## [1] 0.4799297Verifying the previous value:
## [1] 0.4799297Let’s have a look at other goodness-of-fit measures:
mNSE(sim=sim, obs=obs) # Modified NSE## [1] 0.6049584
rNSE(sim=sim, obs=obs) # Relative NSE## [1] -0.5687206
wNSE(sim=sim, obs=obs) # Weighted NSE## [1] 0.9769178
wsNSE(sim=sim, obs=obs) # Weighted Seasonal NSE## [1] 0.9686072
KGE(sim=sim, obs=obs) # Kling-Gupta efficiency (KGE), 2009## [1] 0.6805856
KGE(sim=sim, obs=obs, method="2012") # Kling-Gupta efficiency (KGE), 2012## [1] 0.61689
KGE(sim=sim, obs=obs, method="2021") # Kling-Gupta efficiency (KGE), 2021## [1] 0.7461172
KGElf(sim=sim, obs=obs) # KGE for low flows## [1] 0.5170138
KGEnp(sim=sim, obs=obs) # Non-parametric KGE## [1] 0.6340134
sKGE(sim=sim, obs=obs) # Split KGE## [1] 0.6547522
KGEkm(sim=sim, obs=obs) # Knowable Moments KGE## [1] 0.6467934
JDKGE(sim=sim, obs=obs) # Joint Divergence KGE## [1] 0.6080188
d(sim=sim, obs=obs) # Index of Agreement## [1] 0.9697286
dr(sim=sim, obs=obs) # Refined Index of Agreement## [1] 0.8024792
md(sim=sim, obs=obs) # Modified Index of Agreement## [1] 0.7980307
rd(sim=sim, obs=obs) # Relative Index of Agreement## [1] 0.6231506
VE(sim=sim, obs=obs) # Volumetric Efficiency## [1] 0.6838531
cp(sim=sim, obs=obs) # Coefficient of Persistence## [1] 0.4683536
APFB(sim=sim, obs=obs) # Annual Peak Flow Bias## [1] 0.03202514
HFB(sim=sim, obs=obs) # High Flow Bias## [1] 0.08706977
LME(sim=sim, obs=obs) # Liu-Mean Efficiency## [1] 0.6838398
LCE(sim=sim, obs=obs) # Lee and Choi Efficiency## [1] 0.6777786
PMR(sim=sim, obs=obs) # Proxy for Model Robustness## [1] 0.3156247
pbias(sim=sim, obs=obs) # Percent bias (PBIAS)## [1] 31.6
pbiasfdc(sim=sim, obs=obs) # PBIAS in the slope of the midsegment of the FDC## [Note: 'thr.shw' was set to FALSE to avoid confusing legends...]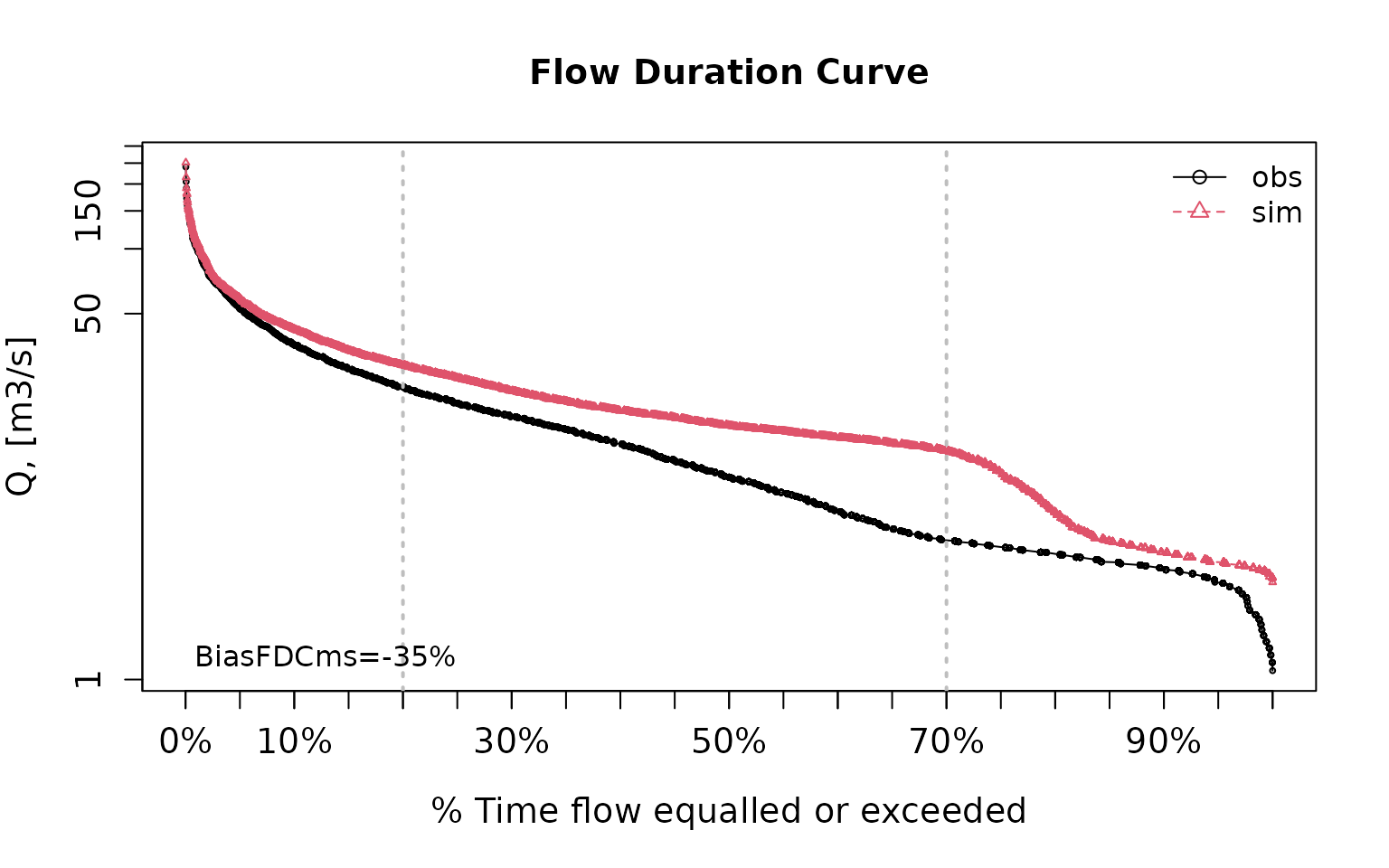
## [1] -34.95419
me(sim=sim, obs=obs) # Mean Error## [1] 4.998998
mae(sim=sim, obs=obs) # Mean Absolute Error## [1] 4.998998
mse(sim=sim, obs=obs) # Mean Squared Error ## [1] 50.46771
rmse(sim=sim, obs=obs) # Root Mean Square Error (RMSE)## [1] 7.104063
ubRMSE(sim=sim, obs=obs) # Unbiased RMSE## [1] 5.047547
nrmse(sim=sim, obs=obs, norm="sd") # Normalised Root Mean Square Error## [1] 35.5
nrmse(sim=sim, obs=obs, norm="maxmin") # Normalised Root Mean Square Error## [1] 3
nrmse(sim=sim, obs=obs, norm="mean") # Normalised Root Mean Square Error## [1] 44.9
nrmse(sim=sim, obs=obs, norm="IQR") # Normalised Root Mean Square Error## [1] 47.1
rPearson(sim=sim, obs=obs) # Pearson correlation coefficient## [1] 0.9698058
rSpearman(sim=sim, obs=obs) # Spearman rank correlation coefficient## [1] 0.8362479
R2(sim=sim, obs=obs) # Coefficient of determination (R2)## [1] 0.8739885
br2(sim=sim, obs=obs) # R2 multiplied by the slope of the regression line## [1] 0.7780545Ex 6: random noise, logarithmic transformation and Pushpalatha2012 constant
NSE for simulated values equal to observations plus random noise on the first half of the observed values and applying (natural) logarithm to obs against sim and adding the Pushpalatha2012 constant during computations.
NSE(sim=sim, obs=obs, fun=log, epsilon.type="Pushpalatha2012")## [1] 0.4871177Verifying the previous value, with the epsilon value following Pushpalatha2012:
## [1] 0.4871177Let’s have a look at other goodness-of-fit measures:
gof(sim=sim, obs=obs, fun=log, epsilon.type="Pushpalatha2012", do.spearman=TRUE, do.pbfdc=TRUE, do.pmr=TRUE)## [,1]
## ME 0.41
## MAE 0.41
## MSE 0.46
## RMSE 0.68
## ubRMSE 0.54
## NRMSE % 71.60
## PBIAS % 18.20
## RSR 0.72
## rSD 0.89
## NSE 0.49
## mNSE 0.48
## rNSE -2.08
## wNSE 0.74
## wsNSE 0.78
## d 0.86
## dr 0.74
## md 0.74
## rd 0.18
## cp -7.68
## r 0.83
## R2 0.49
## bR2 0.44
## VE 0.82
## KGE 0.72
## KGElf 0.52
## KGEnp 0.74
## KGEkm 0.74
## JDKGE 0.72
## LME 0.68
## LCE 0.67
## sKGE 0.53
## APFB 0.01
## HFB 0.02
## rSpearman 0.84
## pbiasFDC % -45.87
## PMR 0.20Ex 7: random noise, logarithmic transformation and user-defined constant
NSE for simulated values equal to observations plus random noise on the first half of the observed values and applying (natural) logarithm to obs against sim and adding a user-defined constant during computations
eps <- 0.01
NSE(sim=sim, obs=obs, fun=log, epsilon.type="otherValue", epsilon.value=eps)## [1] 0.4804Verifying the previous value:
## [1] 0.4804Let’s have a look at other goodness-of-fit measures:
gof(sim=sim, obs=obs, fun=log, epsilon.type="otherValue", epsilon.value=eps, do.spearman=TRUE, do.pbfdc=TRUE, do.pmr=TRUE)## [,1]
## ME 0.42
## MAE 0.42
## MSE 0.48
## RMSE 0.69
## ubRMSE 0.55
## NRMSE % 72.10
## PBIAS % 18.70
## RSR 0.72
## rSD 0.88
## NSE 0.48
## mNSE 0.48
## rNSE -4.22
## wNSE 0.74
## wsNSE 0.78
## d 0.86
## dr 0.74
## md 0.74
## rd -0.39
## cp -7.93
## r 0.82
## R2 0.48
## bR2 0.43
## VE 0.81
## KGE 0.72
## KGElf 0.51
## KGEnp 0.74
## KGEkm 0.73
## JDKGE 0.71
## LME 0.67
## LCE 0.66
## sKGE 0.48
## APFB 0.01
## HFB 0.02
## rSpearman 0.84
## pbiasFDC % -46.36
## PMR 0.21Ex 8: random noise, logarithmic transformation and user-defined factor
NSE for simulated values equal to observations plus random noise on the first half of the observed values and applying (natural) logarithm to obs against sim and using a user-defined factor to multiply the mean of the observed values to obtain the constant to be added to obs against sim during computations
fact <- 1/50
NSE(sim=sim, obs=obs, fun=log, epsilon.type="otherFactor", epsilon.value=fact)## [1] 0.4938294Verifying the previous value:
fact <- 1/50
eps <- fact*mean(obs, na.rm=TRUE)
lsim <- log(sim+eps)
lobs <- log(obs+eps)
NSE(sim=lsim, obs=lobs)## [1] 0.4938294Let’s have a look at other goodness-of-fit measures:
gof(sim=sim, obs=obs, fun=log, epsilon.type="otherFactor", epsilon.value=fact, do.spearman=TRUE, do.pbfdc=TRUE, do.pmr=TRUE)## [,1]
## ME 0.41
## MAE 0.41
## MSE 0.44
## RMSE 0.66
## ubRMSE 0.52
## NRMSE % 71.10
## PBIAS % 17.60
## RSR 0.71
## rSD 0.89
## NSE 0.49
## mNSE 0.48
## rNSE -1.33
## wNSE 0.74
## wsNSE 0.78
## d 0.87
## dr 0.74
## md 0.74
## rd 0.38
## cp -7.43
## r 0.83
## R2 0.49
## bR2 0.44
## VE 0.82
## KGE 0.73
## KGElf 0.53
## KGEnp 0.74
## KGEkm 0.75
## JDKGE 0.72
## LME 0.68
## LCE 0.68
## sKGE 0.56
## APFB 0.01
## HFB 0.02
## rSpearman 0.84
## pbiasFDC % -45.37
## PMR 0.19Ex 9: random noise, user-defined transformation
NSE for simulated values equal to observations plus random noise on the first half of the observed values and applying a user-defined function to obs against sim during computations:
## [1] 0.7265255Verifying the previous value, with the epsilon value following Pushpalatha2012:
## [1] 0.7265255
gof(sim=sim, obs=obs, fun=fun1, do.spearman=TRUE, do.pbfdc=TRUE, do.pmr=TRUE)## [,1]
## ME 0.65
## MAE 0.65
## MSE 0.92
## RMSE 0.96
## ubRMSE 0.71
## NRMSE % 52.30
## PBIAS % 17.70
## RSR 0.52
## rSD 0.97
## NSE 0.73
## mNSE 0.54
## rNSE 0.34
## wNSE 0.89
## wsNSE 0.76
## d 0.93
## dr 0.77
## md 0.76
## rd 0.83
## cp -1.17
## r 0.92
## R2 0.73
## bR2 0.65
## VE 0.82
## KGE 0.81
## KGElf 0.50
## KGEnp 0.75
## KGEkm 0.80
## JDKGE 0.79
## LME 0.80
## LCE 0.79
## sKGE 0.84
## APFB 0.02
## HFB 0.04
## rSpearman 0.84
## pbiasFDC % -32.96
## PMR 0.19A short example from hydrological modelling
Loading observed streamflows of the Ega River (Spain), with daily data from 1961-Jan-01 up to 1970-Dec-31
Generating a simulated daily time series, initially equal to the observed values (simulated values are usually read from the output files of the hydrological model)
sim <- obs Plotting the graphical comparison of obs against sim, along with the default goodness-of-fit measures:
ggof(sim, obs)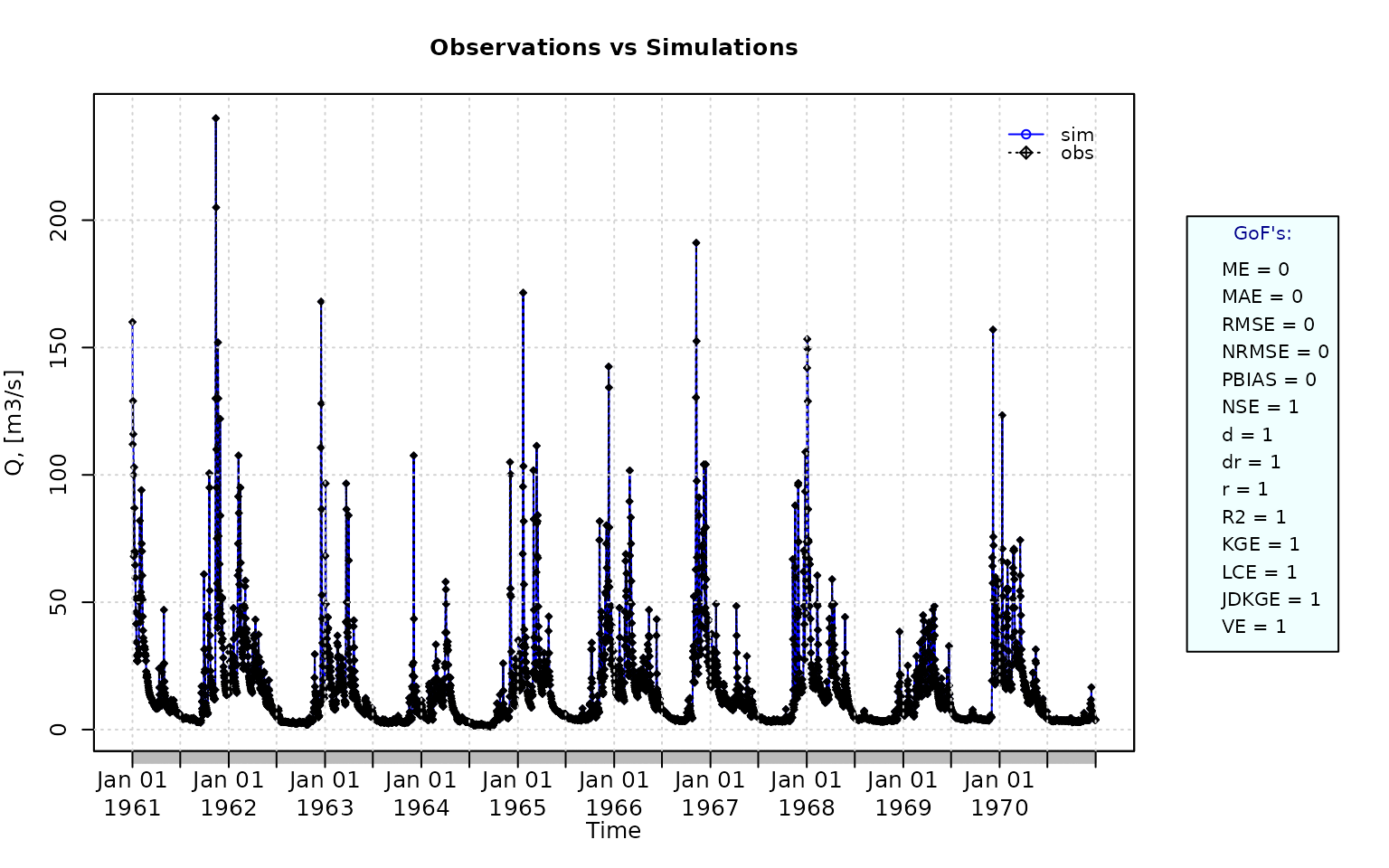
Plotting the graphical comparison of obs against sim, along with user-defined goodness-of-fit measures:
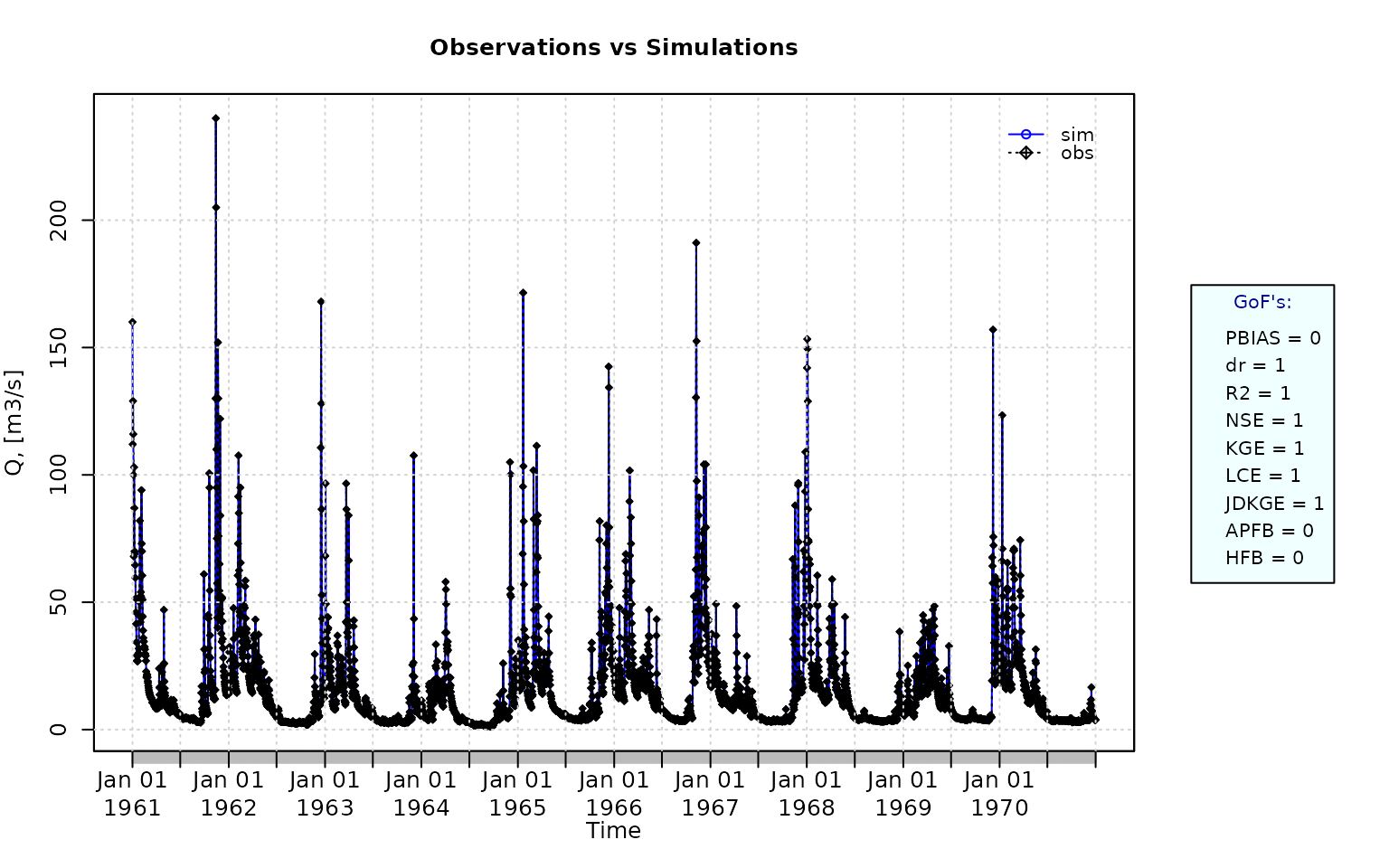
Computing the numeric goodness-of-fit measures for the best (unattainable) case, including the following three measures that -due to slightly high computation time- are not computed by default: i) the Spearman’s Rank Correlation Coefficient (rSpearman), ii) the percent bias in the Slope of the Midsegment of the Flow Duration Curve (pbiasFDC), and iii) the Proxy for Model Robustness (PMR).
gof(sim=sim, obs=obs, do.spearman=TRUE, do.pbfdc=TRUE, do.pmr=TRUE)## [,1]
## ME 0
## MAE 0
## MSE 0
## RMSE 0
## ubRMSE 0
## NRMSE % 0
## PBIAS % 0
## RSR 0
## rSD 1
## NSE 1
## mNSE 1
## rNSE 1
## wNSE 1
## wsNSE 1
## d 1
## dr 1
## md 1
## rd 1
## cp 1
## r 1
## R2 1
## bR2 1
## VE 1
## KGE 1
## KGElf 1
## KGEnp 1
## KGEkm 1
## JDKGE 1
## LME 1
## LCE 1
## sKGE 1
## APFB 0
## HFB 0
## rSpearman 1
## pbiasFDC % 0
## PMR 0- Randomly changing the first 1826 elements of ‘sim’ (half of the ts), by using a normal distribution with mean 10 and standard deviation equal to 1 (default of ‘rnorm’).
sim[1:1826] <- obs[1:1826] + rnorm(1826, mean=10)Plotting the graphical comparison of obs against sim, along with the default goodness-of-fit measures for the daily and monthly time series:
ggof(sim=sim, obs=obs, ftype="dm", FUN=mean)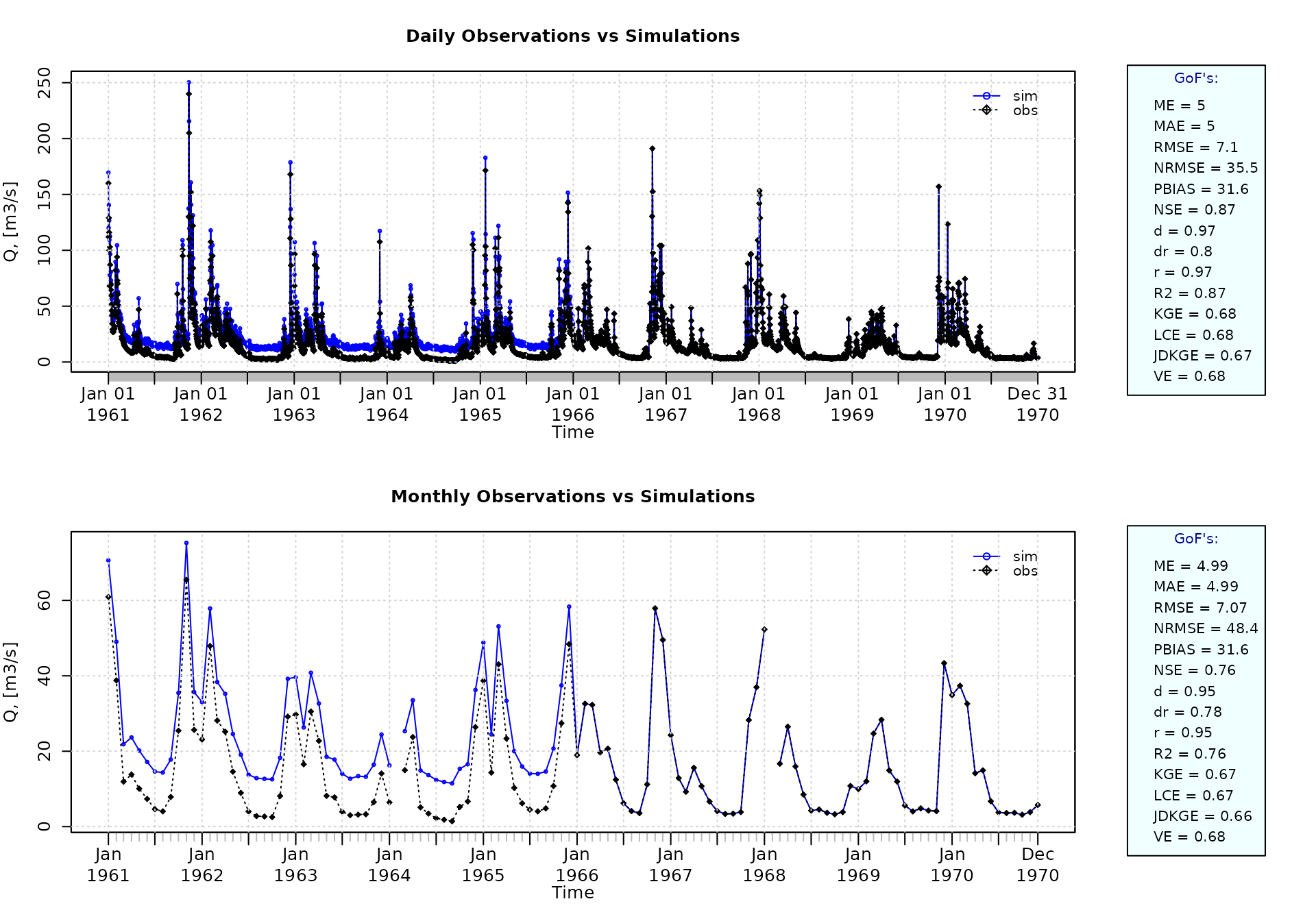
Plotting the graphical comparison of obs against sim, along with user-defined goodness-of-fit measures for the daily and monthly time series:
ggof(sim=sim, obs=obs, ftype="dm", FUN=mean,
gofs=c( "PBIAS", "dr", "R2", "NSE", "KGE", "LCE", "JDKGE", "APFB", "HFB"))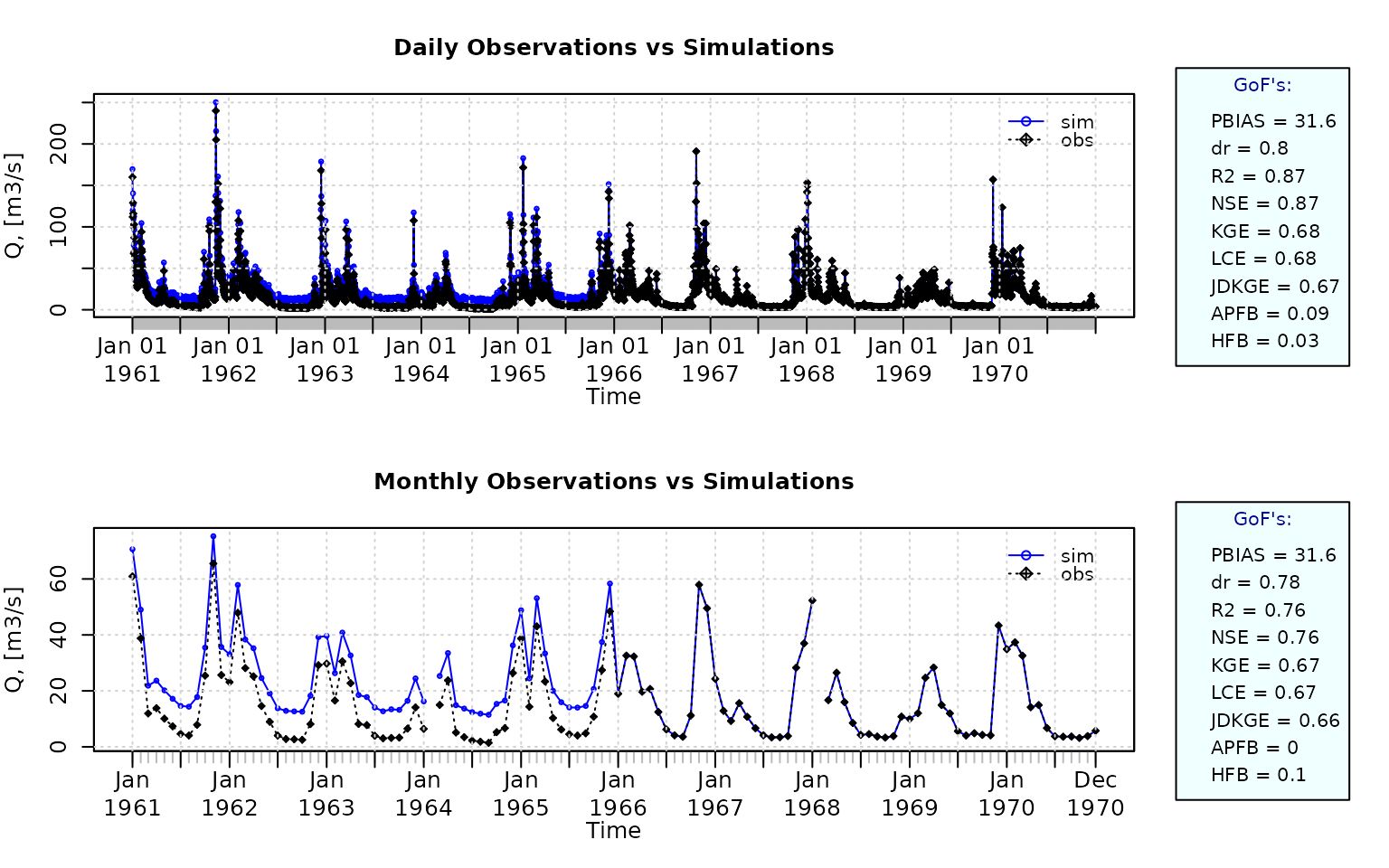
Removing warm-up period
Using the first two years (1961-1962) as warm-up period, and removing the corresponding observed and simulated values from the computation of the goodness-of-fit measures:
ggof(sim=sim, obs=obs, ftype="dm", FUN=mean, cal.ini="1963-01-01")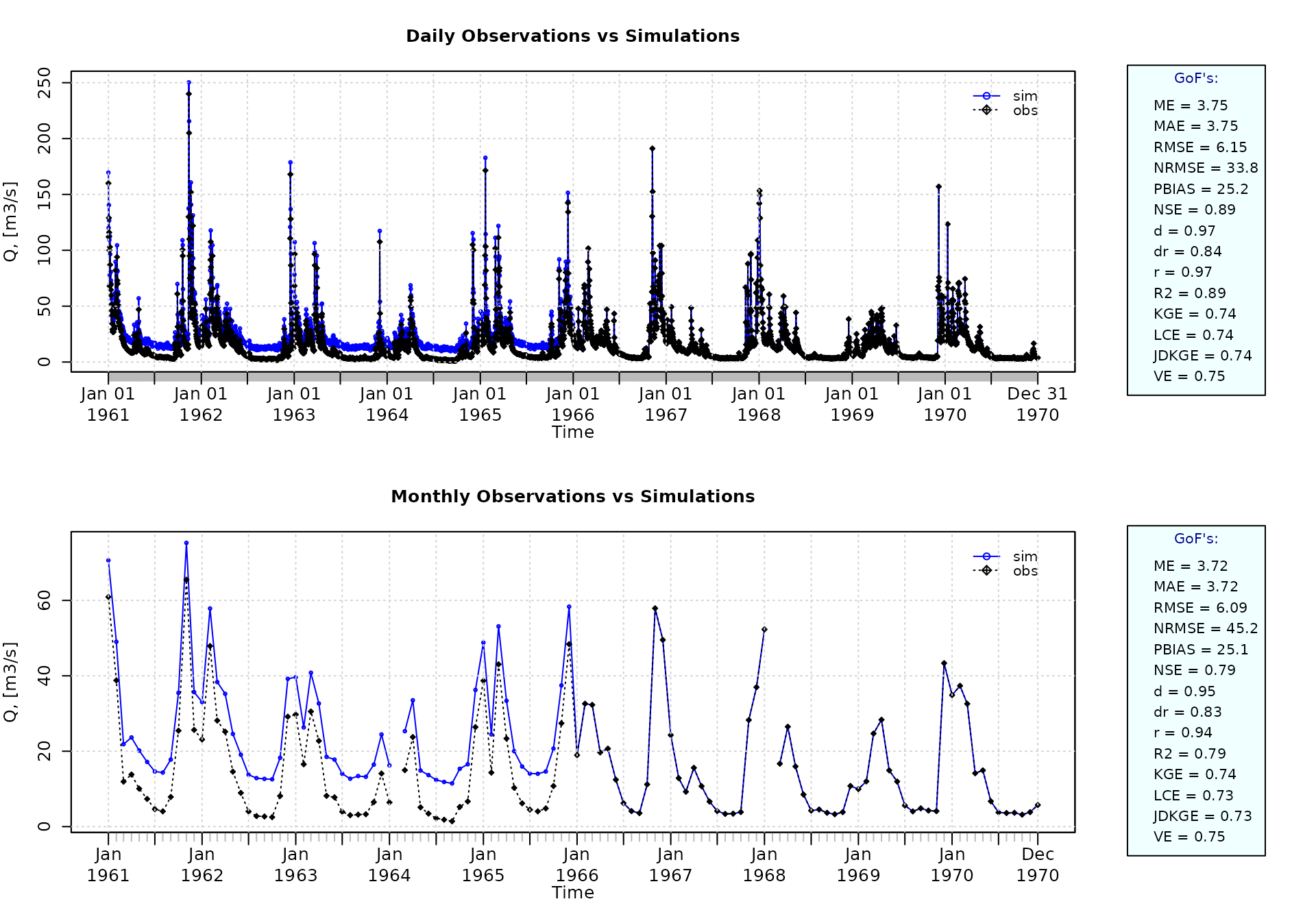
Verification of the goodness-of-fit measures for the daily values after removing the warm-up period:
## [,1]
## ME 3.75
## MAE 3.75
## MSE 37.84
## RMSE 6.15
## ubRMSE 4.88
## NRMSE % 33.80
## PBIAS % 25.20
## RSR 0.34
## rSD 1.02
## NSE 0.89
## mNSE 0.68
## rNSE -0.48
## wNSE 0.98
## wsNSE 0.82
## d 0.97
## dr 0.84
## md 0.84
## rd 0.64
## cp 0.52
## r 0.97
## R2 0.89
## bR2 0.81
## VE 0.75
## KGE 0.74
## KGElf 0.57
## KGEnp 0.69
## KGEkm 0.73
## JDKGE 0.74
## LME 0.75
## LCE 0.74
## sKGE 0.70
## APFB 0.03
## HFB 0.00Plotting uncertainty bands
Generating fictitious lower and upper uncertainty bounds:
lband <- obs - 5
uband <- obs + 5Plotting the previously generated uncertainty bands:
plotbands(obs, lband, uband)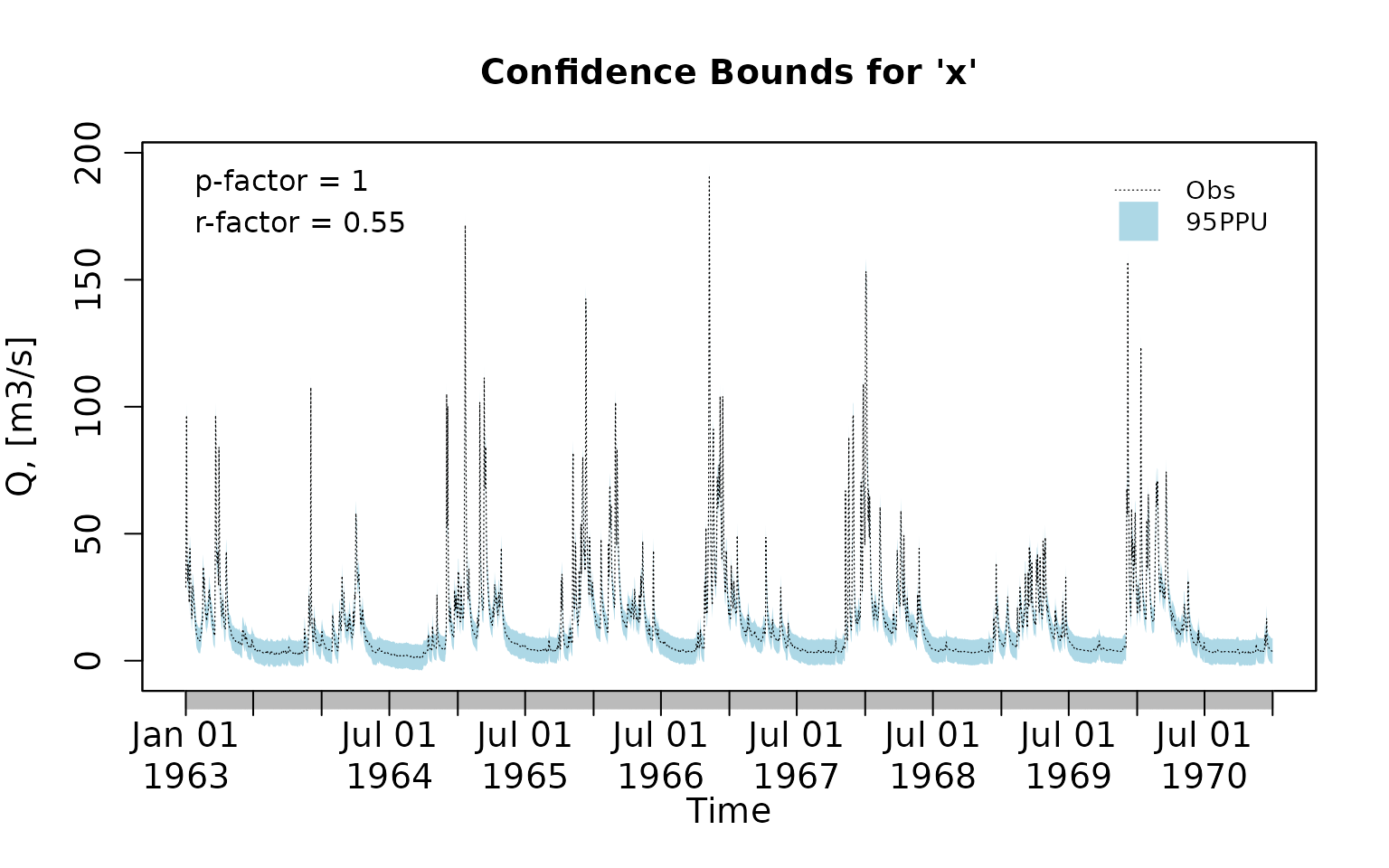
Randomly generating a simulated time series:
Plotting the previously generated simulated time series along the observations and the uncertainty bounds:
plotbands(obs, lband, uband, sim)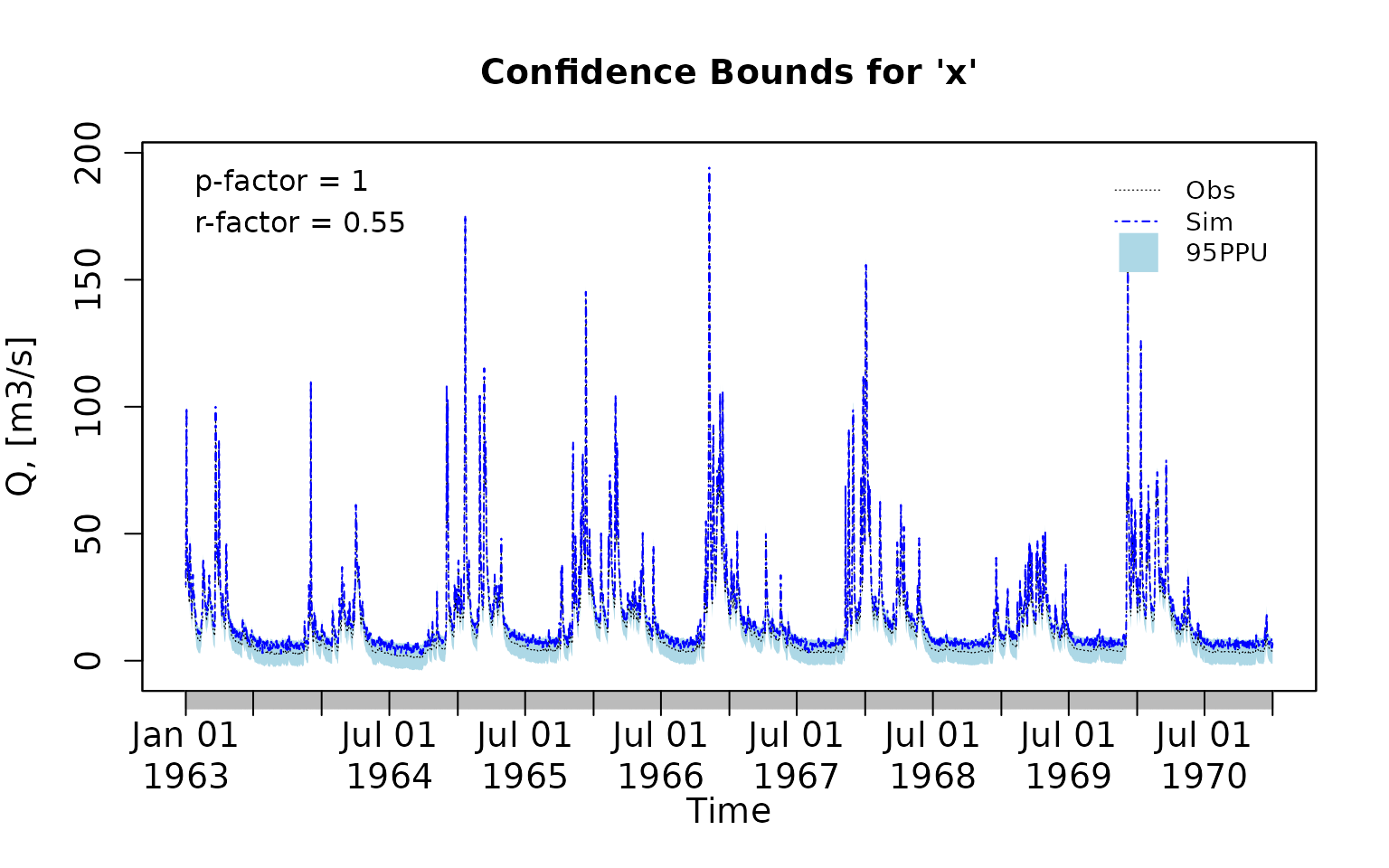
Computing P-factor
The P-factor quantifies the percentage of observed values that fall within the prediction uncertainty band defined by the lower and upper bounds. It is a measure of the coverage of the uncertainty interval.
The P-factor ranges from 0 to 1. A value of 1 indicates that all observations are bracketed by the uncertainty bounds, whereas a value of 0 indicates that none of the observations fall within the bounds.
Computing the P-factor:
( pfactor(x=obs, lband=lband, uband=uband) )## [1] 0.9993155Computing R-factor
The R-factor quantifies the average width of the prediction uncertainty band relative to the variability of the observed data. It is a measure of the magnitude of predictive uncertainty associated with a model simulation.
The R-factor ranges from 0 to infinity, with an optimal value of 0 indicating perfect agreement between simulated and observed values (i.e., zero prediction uncertainty). In practical applications, the R-factor represents the width of the uncertainty interval and should be as small as possible. Values close to or smaller than 1 are commonly considered indicative of an acceptable level of predictive uncertainty, although acceptable thresholds depend on the quality of observations and the modelling context.
Computing the R-factor:
( rfactor(x=obs, lband=lband, uband=uband) )## [1] 0.5488274Important note:
Because a larger fraction of observations can often be bracketed by widening the uncertainty bounds, the R-factor is typically interpreted jointly with the P-factor. A balance between a high P-factor (good coverage) and a low R-factor (narrow uncertainty bounds) is therefore sought during model calibration and uncertainty analysis.
Analysis of the residuals
Computing the daily residuals (even if this is a dummy example, it is enough for illustrating the capability)
r <- sim-obsSummarizing and plotting the residuals (it requires the hydroTSM package):
## Index r
## Min. 1963-01-01 -0.3042
## 1st Qu. 1964-12-31 2.3620
## Median 1966-12-31 3.0400
## Mean 1966-12-31 3.0310
## 3rd Qu. 1968-12-30 3.7110
## Max. 1970-12-31 6.4810
## IQR <NA> 1.3488
## sd <NA> 1.0070
## cv <NA> 0.3323
## Skewness <NA> -0.0700
## Kurtosis <NA> -0.0253
## NA's <NA> 2.0000
## n <NA> 2922.0000
# daily, monthly and annual plots, boxplots and histograms
hydroplot(r, FUN=mean)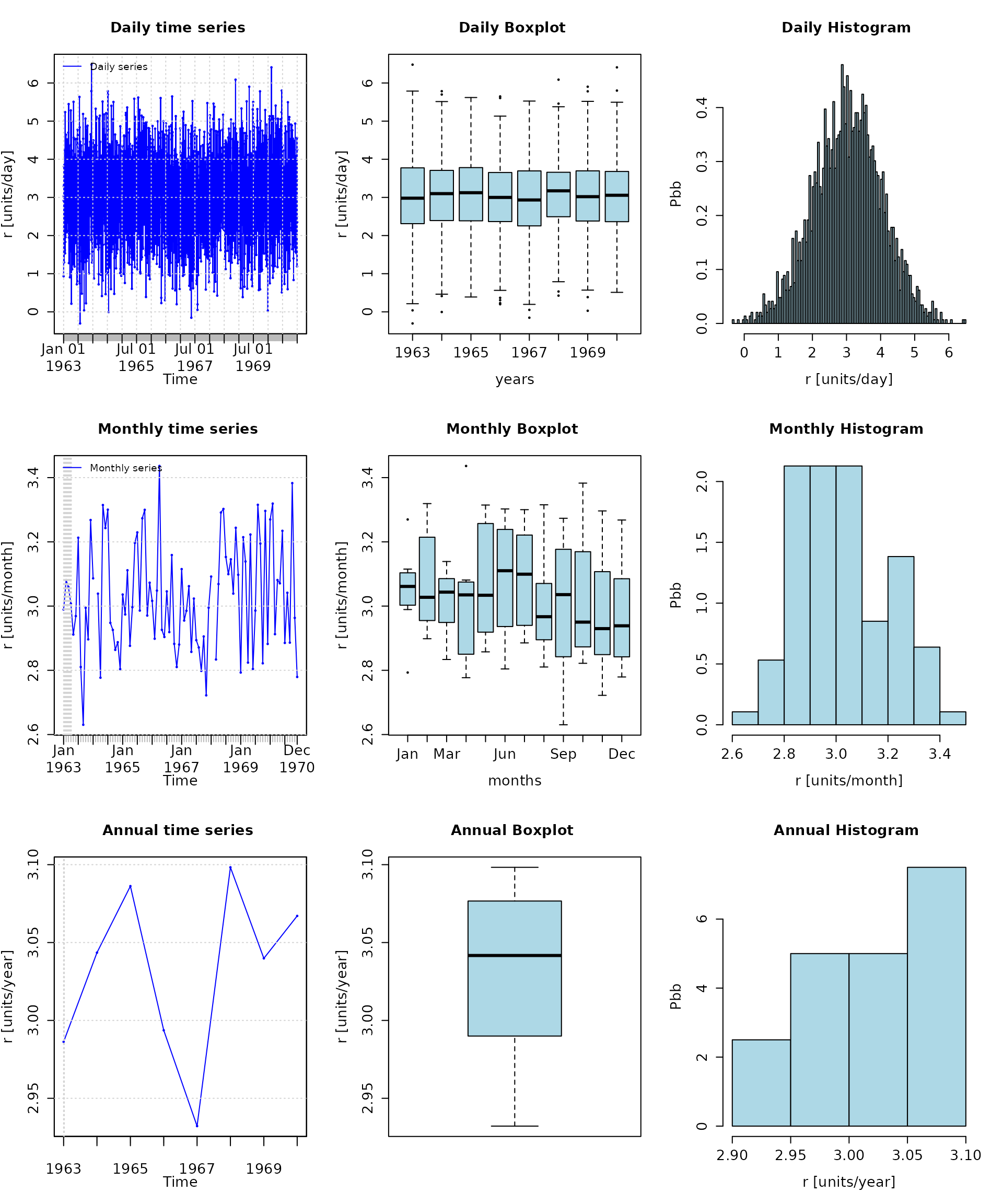
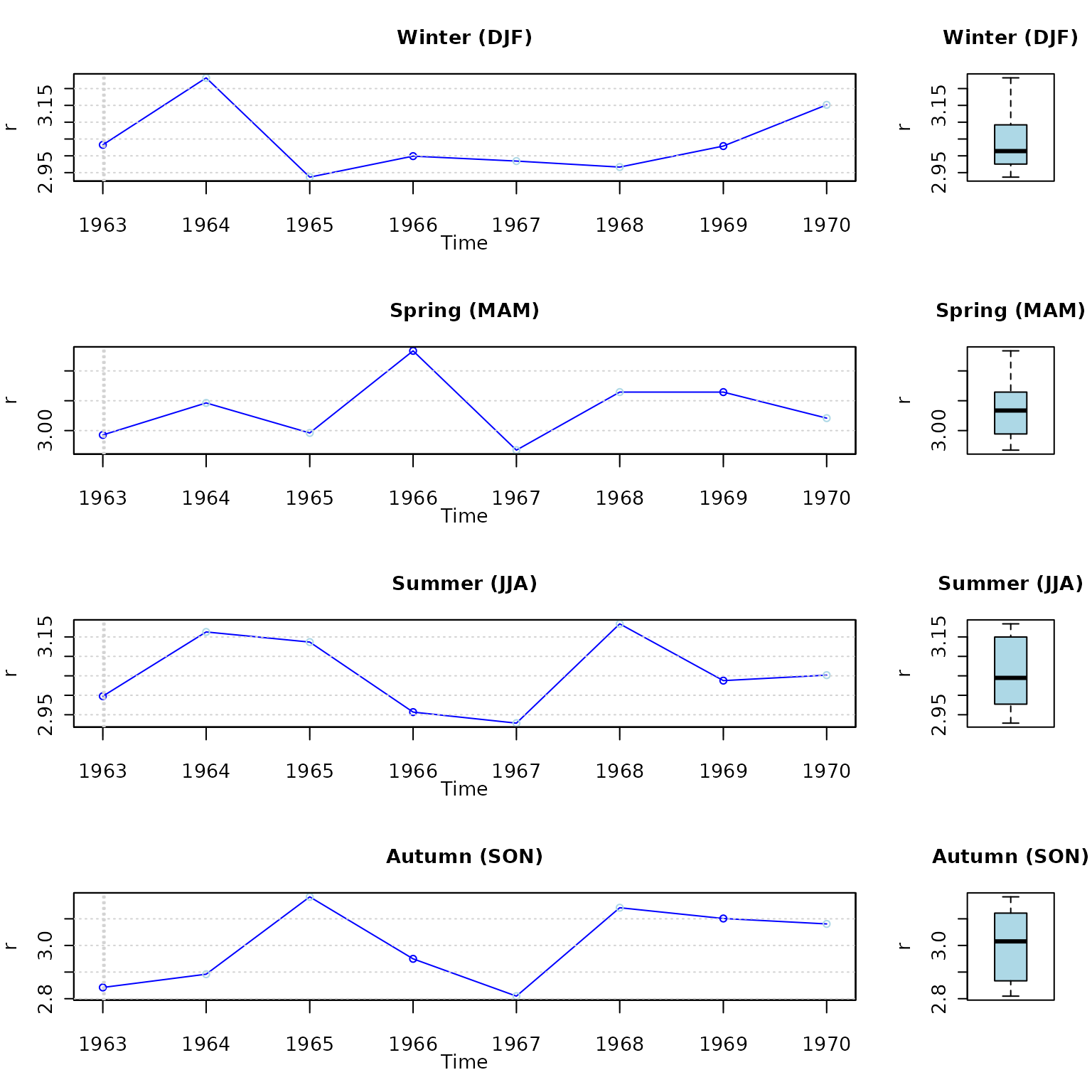
Seasonal plots and boxplots
# daily, monthly and annual plots, boxplots and histograms
hydroplot(r, FUN=mean, pfreq="seasonal")